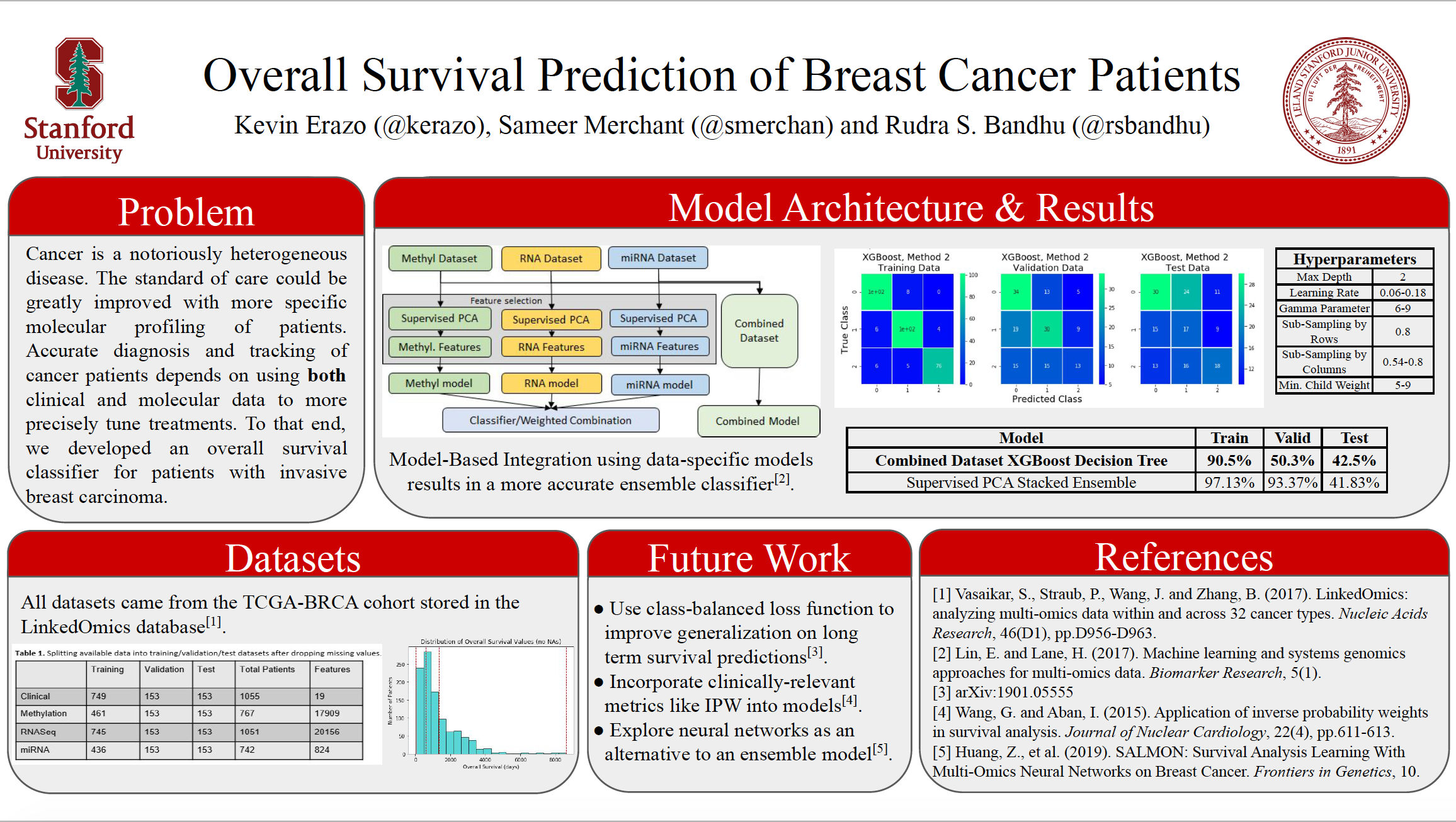
Combining Multi-Omics Data to Predict Overall Survival of Breast Cancer Patients
Precision medicine, an emerging field of medicine, has enabled prospects of customization ofhealthcare, medical decisions and treatments tailored to individual patients.The use of genomic andtranscriptomic biomarkers as well as other multi-omics data has played a major role in precision oncology.Concurrent with the explosion of clinically relevant molecular data, the application of machine learningmethods to multi-omics datasets has become more commonplace. In this paper we present a novel methodof combining multi-omics datasets to predict breast cancer patients’ overall survival
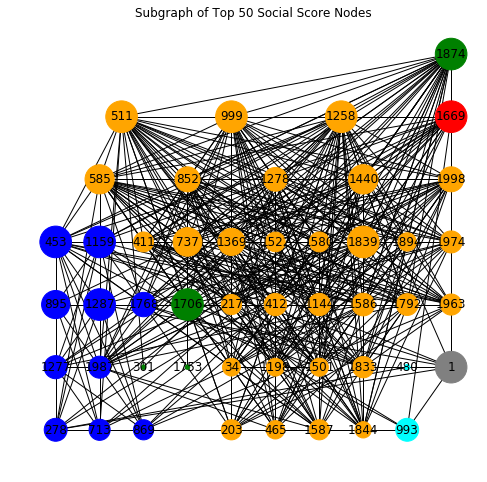
Analyzing and Mitigating Phishing Outbreaks: a Case Study on the 2016 DNC Email Network
In this project, we take a perspective of a system administrator seeking to secure an email network. Given budget constraint in the form of number of detectors (cybersecurity resources) available, how do we secure our email network by maximizing likelihood of outbreak detection, minimizing detection time and reducing the population affected? This project aims to optimally detect phishing outbreaks in an emperical email network dataset that maps the communications of 2016 Democratic National Committee. We use CELF as a benchmark for choosing "detector" nodes in our outbreak. While CELF is very effective in placing detector nodes, it can be computationally expensive and its runtime can be prohibitive when simulating large numbers of outbreaks in a dynamically changing email network. We investigate methods of speeding up CELF through use of a node metric called Social Score, to assist CELF with node selection. Our experimental results on the DNC email network shows a 10x speed up in selecting detector nodes. Based on our results, we believe Social Score serves as an effective heuristic to speed up CELF despite expected decreases in quality of selection.
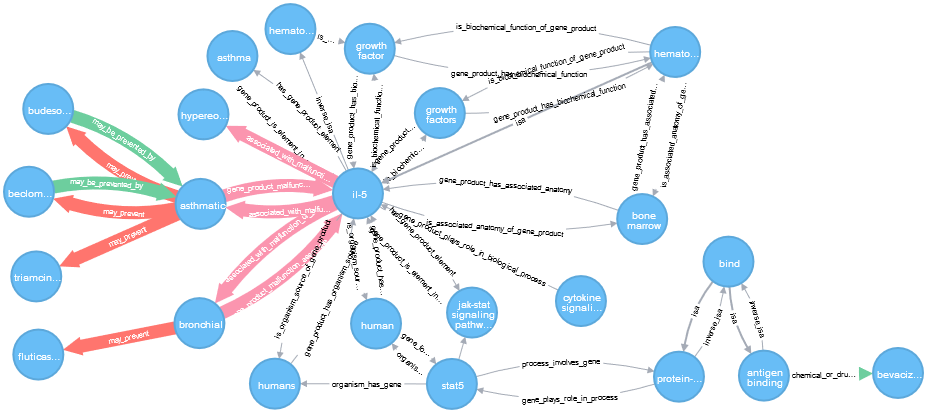
Predicting Disease Treatments from Biomedical Knowledge Graph Data
Prescribing drugs for off-label uses is a common practice in medicine. Drugs have been repositioned via mining clinical data, applying statistical tools, and leveraging molecular information. Computational analysis of biomedical knowledge allows us to discover new uses for existing drugs on the market. In this report, we focus on the problem of predicting whether a given drug can treat a given disease. We examined different mechanisms of creating a metapath-based models to predict drug-disease relations using information encoded in a biomedical knowledge graph. Our proposed metapath-based model achieved AUROC=0.912 in 10-fold cross validation.